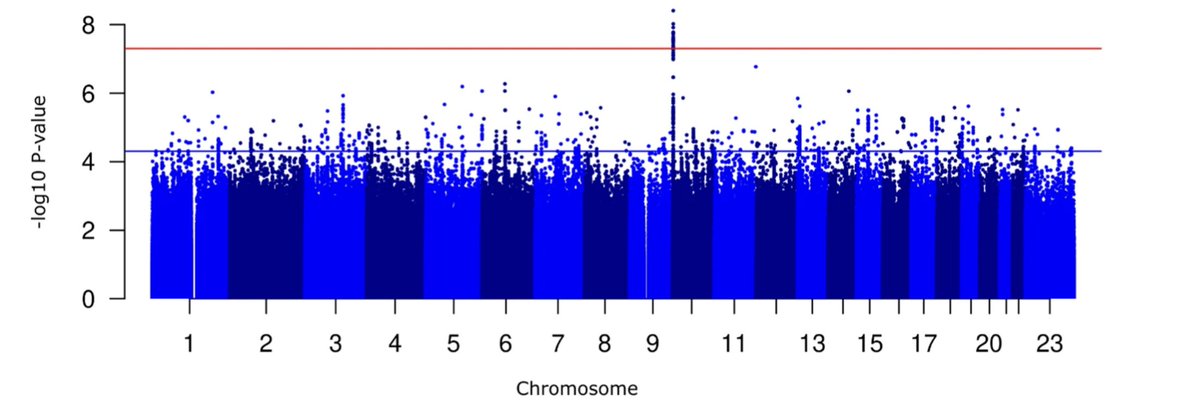
Last but not least, Sam Deakin presented some his PhD work in the Coltman Lab on QTL mapping (GWAS) for fitness-related traits in Ram Mountain bighorn sheep and possible impacts of trophy hunting on QTL allele frequencies #CSEE2024



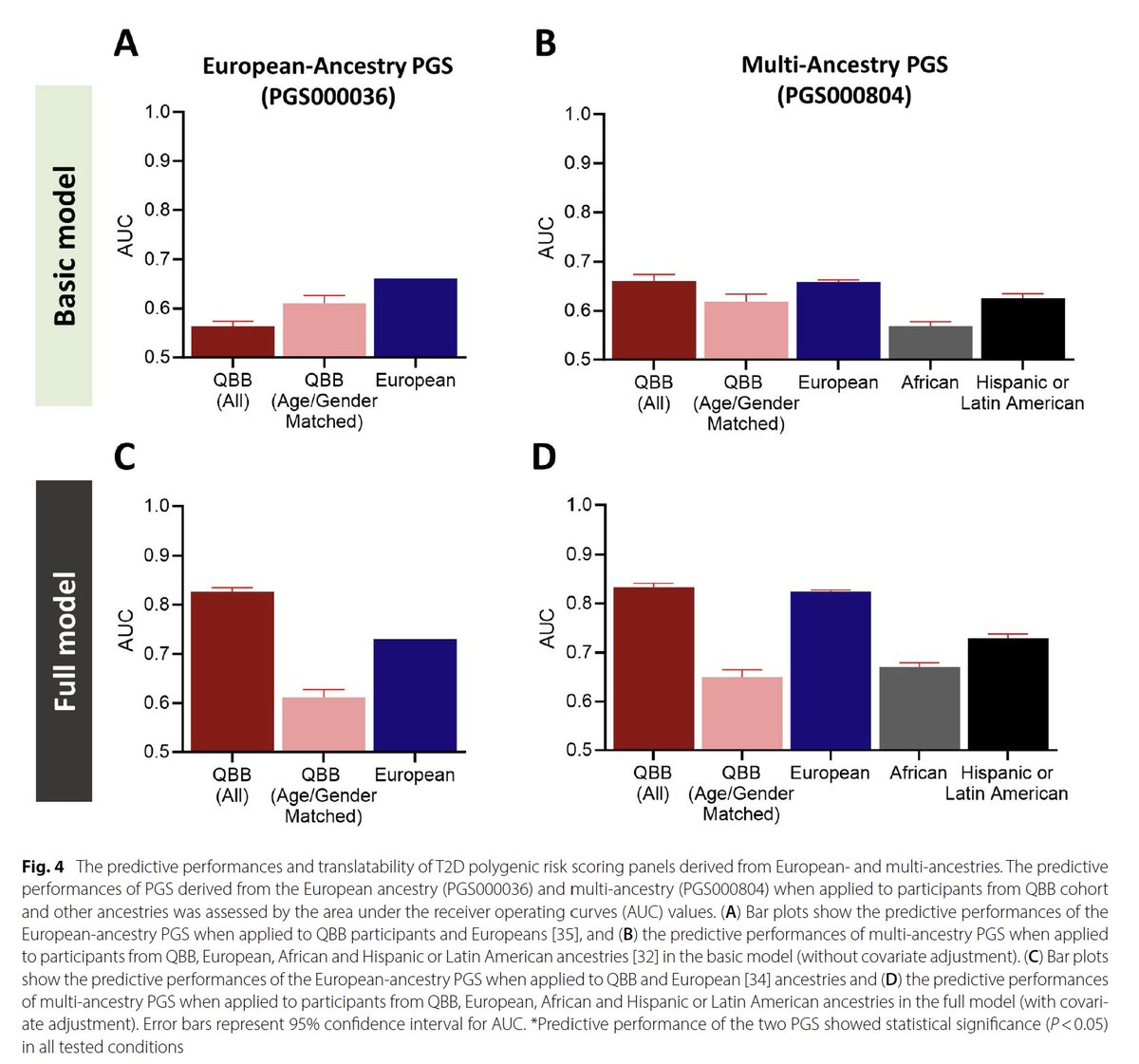
Here the latest #GWAS study with #T2D in Qatar Biobank, led by Omar Albagha from جامعة حمد بن خليفة :
Genome-wide association study and trans-ethnic meta-analysis identify novel susceptibility loci for type 2 #diabetes mellitus.


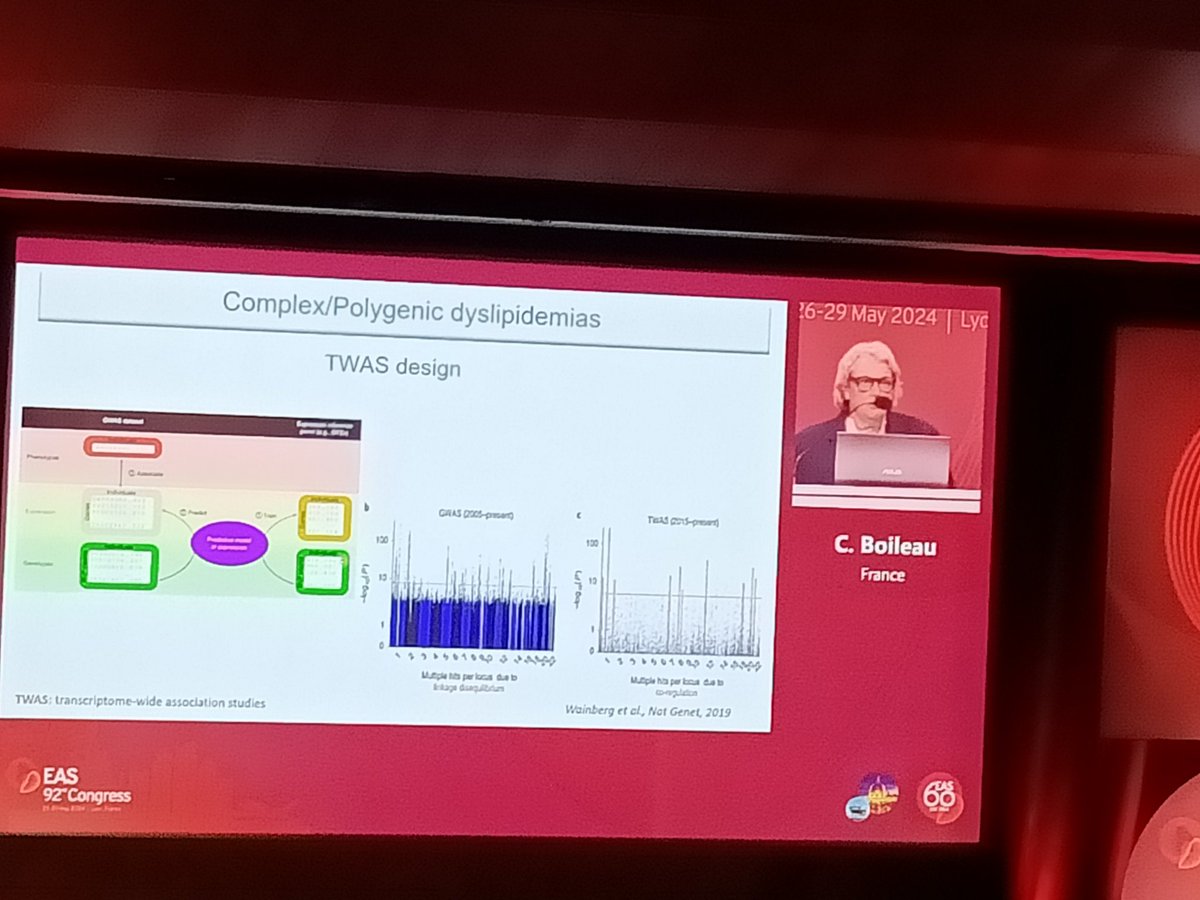
From #GWAS to #TWAS to better disentangle the complex background underneath the most-difficult-to-understand #dyslipidemias and go to the ♥️ of the problem!
#EASCongress2024
EAS Congress European Atherosclerosis Society
HELP! Do you use GWAS Catalog data and/or any other data managed by EMBL-EBI?
✔️ Fill out the survey shorturl.at/wOCO3
🕰️ Less than 10 min
🗓️ Before 7th June
#openaccess





Very excited that my meta-analysis of telomere length GWAS and experimental validation showing that POP5 and KBTBD6 are human telomere length regulation genes is out in Nature Communications!
Shout out to my mentors Alexis Battle and Rasika Mathias
nature.com/articles/s4146…

Day 1 of 'A Hitchhiker's Guide to Statistical Genetics Principles and Practice' course: LD, h2, genetic models, GWAS. Nicola Pirastu Health Data Science Centre at Human Technopole Sodbo Sharapov 🌌 We had uniforms.


.Kaikkonen Lab giving the keynote talk at #VascularDiscovery24 on functional annotation of Coronary Artery Disease GWAS loci.






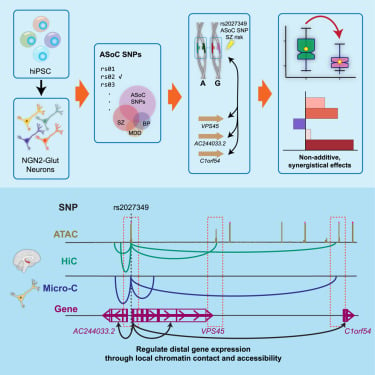
Multiple genes in a single GWAS risk locus synergistically mediate aberrant synaptic development and function in human neurons
hubs.li/Q02v7f5C0
Up-and-coming genomics research from Cell Genomics #neurogenomics